HepaFat-Scan
MRI Measurement Of Liver Fat
HepaFat-Scan is an FDA cleared MRI-based solution for measuring hepatic steatosis in patients at risk from fatty liver disease. It provides a reliable non-invasive measure of the volume fraction of fat in liver tissue for screening, diagnosis, treatment planning and monitoring of patients in a variety of clinical settings.
The HepaFat-Scan service is a 2-minute MRI scan followed by off-site processing of the images in Resonance Health’s ISO certified central image analysis centre. These images, acquirable on most makes and models of 1.5 and 3 Tesla MRI machines, are obtained through a unique and standardised scanning sequence to ensure results are accurate, reliable, and reproducible over time and between hospitals and the various makes and models of MRI scanners.
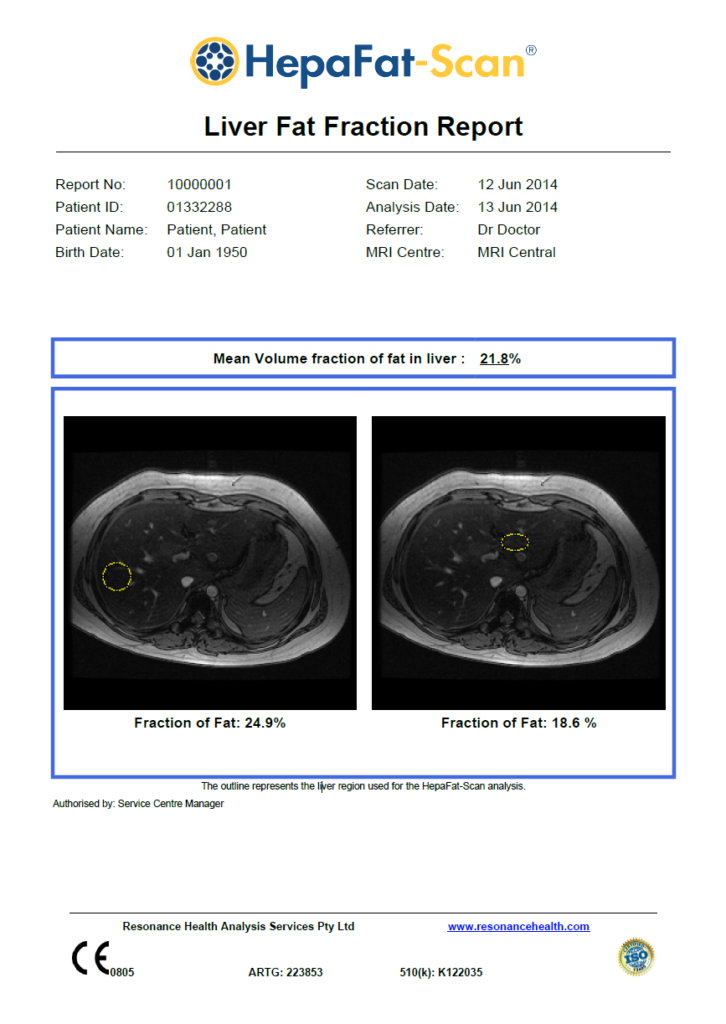
The HepaFat-Scan Report
HepaFat-Scan reports the volumetric fraction of fat in the liver. The fractional area of a liver biopsy histological section that appears as fat is equivalent to the volumetric liver fat fraction and thus can be compared directly with HepaFat-Scan results.
Other MRI methods for assessing liver steatosis are either semi-quantitative or report the ratio of protons in fat and water molecules (referred to as the Proton Density Fat Fraction).
HepaFat-Scan is available for routine clinical use, investigator led clinical research and drug development.
Key HepaFat-Scan Features
- Non-invasive, contrast free with a short scan time of only two minutes with a single breath hold
- Results are available within two (2) business days
- Suitable for 1.5 and 3 Tesla MRI scanners
- No need to purchase expensive new software or hardware
- International regulatory clearances (USA, Europe, Australia)
- High sensitivity and specificity when compared against histopathological assessment of steatosis
- Accurate, reliable and reproducible over time and between MRI centres and scanner types, it is ideally suited to multicenter clinical trials
- Centralised image analysis and reporting within our ISO 13485 certified Service Centre
- Clinically validated against biopsy (n=59 cases) with no statistically significant bias (1.4%)
- A large liver region analysed >300 times larger than biopsy, meaning sampling errors are minimised
Potential Clinical Applications
Gastroenterology and Hepatology Applications
- Initial workup for Non-alcoholic Fatty Liver Disease (NAFLD) diagnosis and for education and counselling of metabolic syndrome patients.
- Liver fat analysis on patients already being screened or monitored for fibrosis or cirrhosis. Recent research indicates that patients with NAFLD and Non-Alcoholic Steatohepatitis (NASH) can develop liver cancer without progressing through cirrhosis and that higher liver fat in NAFLD is associated with fibrosis progression.
- Screening and monitoring of participants for phase I and II pharmaceutical trials of compounds to treat NASH or diabetes.
Surgical Applications
- Pre and post-operative analysis of bariatric patients to track clinical outcomes. Bariatric patients post-surgery can see a dramatic improvement in liver fat regardless of their starting anthropometrics (e.g. weight, BMI,) or amount of weight loss following surgery.
- A high level of liver fat may have a detrimental outcome on liver surgery. A HepaFat-Scan of patients with liver disease or liver cancer can provide guidance with treatment planning. A quantitative measurement of liver fat may help to determine how much liver can be safely resected or to determine whether to use a surgical or non-surgical treatment.
- Screening for suitability of living donors for liver transplants by assisting in determining the viability of the donor liver.
Applications by Primary Care Physicians
- HepaFat-Scan may be useful to screen patients prior to prescribing known hepatotoxic medications.
- Monitoring of patients undergoing an intervention (e.g. limiting alcohol consumption or weight loss).
- Monitoring of patients prescribed medications known to induce steatosis.
Why Use HepaFat-Scan to Measure Liver Fat
Histopathological assessment of a patient’s liver fat levels from a liver biopsy is commonly considered the gold standard for clinical assessment of liver fat. However, it is subjective, has poor reproducibility, significant sampling errors and suffers from the health risks associated with an invasive procedure.
Ultrasound, though widely available, has poor sensitivity (approximately 70%) and is further reduced in obese patients and those with fibrosis, edema and extrahepatic adipose tissue.
CT is also a widely available imaging modality that evaluates hepatic steatosis indirectly based on hepatic x-ray attenuation. Several factors other than fat (e.g. iron, copper, glycogen, fibrosis, edema, ingestion of drugs such as amiodarone and gold) affect CT attenuation values, resulting in unavoidable errors in fat quantification and low sensitivity for mild-moderate steatosis. Moreover, as CT relies on ionizing radiation, it is not suitable for use in children and for longitudinal monitoring of adults with liver fat. MR has also been used for quantitative assessment of fat and is widely accepted as superior compared to ultrasound and CT.
HepaFat-Scan operates independently from MRI scanners. It is used in conjunction with a specific MRI data acquisition protocol that is a variation of the three point Dixon Method. Image data acquisition for a HepaFat-Scan takes a single breath hold and no contrast agents are required. Critically, and unlike Proton Density Fat Fraction (PDFF) methods, a high flip angle is used to enhance signal to noise ratios (SNR) and hence improve sensitivity and specificity of fat measurements. The SNR of HepaFat-Scan can be between 60% to 82% higher in comparison to a standard PDFF method. Our unique acquisition approach means that HepaFat-Scan is better at discriminating small changes in the fraction of tissue protons in fat, making it ideal for longitudinal monitoring.
When comparing fat measurements obtained from biopsy with measurements obtained using Magnetic Resonance (MR) it is important to ensure the measures are directly comparable. Regular histopathological assessment of liver fat according to the NASH CRN grading scheme does not normally allow direct comparison to an MR measurement.
Other MR fat quantification methods such as Magnetic Resonance Spectroscopy (MRS) and PDFF based imaging methods report the fat : water proton ratio. This ratio is not directly comparable to a fat measurement from a liver biopsy specimen as it neither represents the percentage of parenchyma involved with fat nor the volume fraction of fat in the biopsy. Furthermore, as the proton density in fat is not equal to the proton density of liver tissue there is a non-linear relationship between the liver fat volume fraction and the fat : water proton ratio. Hence, a 5 percent change in fat : water proton ratio does not equate to a 5% change of fat in the liver.
Resonance Health has addressed this issue with HepaFat-Scan. The HepaFat-Scan method takes into account T2* signal decay, the difference in T1 between liver tissue and triglycerides and the densities of protons in triglycerides and liver tissue. By taking into account the difference in proton density between fat and liver tissue, HepaFat-Scan is able to report the percentage volume within the analysed liver region that is fat. This is a measure that is directly comparable to what can be seen and measured in a biopsy specimen. The biopsy samples of the 59 subjects in the clinical study were all assessed using a technique called stereology, which is a computer assisted analysis of each biopsy specimen to quantify the volume fraction of the biopsy that is fat. Hence, the HepaFat-Scan results can be directly compared against the biopsy measurements as both report a volume percentage of fat.
In addition to these advantages, HepaFat-Scan delivers a consistent, standardised image acquisition and analysis service for the measurement of liver fat. In comparison, there are multitudes of PDFF acquisition and processing methods that can make comparisons across studies tenuous. For example, MRI-PDFF data has been acquired with flip angles between 2 to 20 degrees and TR’s between 8 and 200 ms, with 3 to 16 echoes. Processing can use magnitude data or magnitude and complex images, single to multiple peak fitting, and single or multiple T2* fitting. The range of acquisition and processing options used to generate PDFF data can potentially lead to inconsistencies between PDFF studies. For example, the PDFF cut off used to separate grade 0 from grade 1 hepatic steatosis is typically set at 5% fat fraction. However, studies have reported PDFF cut-offs between 1.4 and 17% for discriminating grade 0 from grade 1.
With so many variations of PDFF available, choosing HepaFat-Scan stands out as a consistent, sensitive, and accurate measure of liver steatosis that is directly relatable to biopsy measurements universally trusted across the clinical community.

HepaFat-Scan Clinical Validation
Resonance Health has completed a clinical study comparing 59 patient measurements of volumetric liver fat fraction using HepaFat-Scan and stereological analysis of liver biopsy sections. Stereology is a computer assisted analysis of each biopsy specimen to quantify the volume fraction of the biopsy that is fat. The HepaFat-Scan results can be directly compared against the biopsy measurements as both report a volume percentage of fat. The results of the comparison are illustrated in the figure below. A linear relationship between the two sets of measurements was found with a clinically insignificant bias (1.4% by Bland Altman analysis) between the two types of measurement.
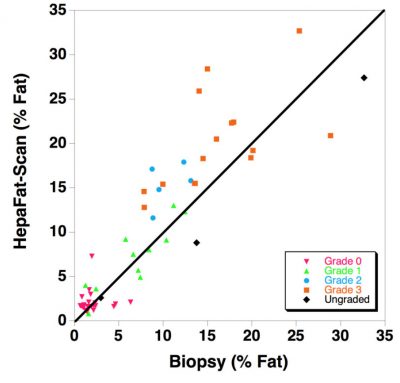
The figure displays a comparison of HepaFat-Scan measurements of liver fat fraction with those measured from biopsy sections using stereology (n=59). The solid line is the line of equivalence. Symbols are colour/shape-coded to indicate the Non-alcoholic Steatohepatitis Clinical Research Network (NASH CRN) steatosis grade assessed by a histopathologist (See below table) [Design and Validation of a Histological Scoring System for Non-alcoholic Fatty Liver Disease, Hepatology, June 2005, Kleiner et al]:
- Grade 0: < 5% parenchymal involvement by steatosis
- Grade 1: 5% - 33% parenchymal involvement by steatosis
- Grade 2: >33%-66% parenchymal involvement by steatosis
- Grade 3: >66% parenchymal involvement by steatosis
All grades of steatosis were represented in the study [Grade 0 (47%); Grade 1 (13%); Grade 2 (9%); Grade 3 (30%)]. The patients in the study had a range of aetiologies including autoimmune hepatitis (n=3), alcoholic liver disease (n=2), chronic viral hepatitis B/C (n=16), non-alcoholic fatty liver disease (n=10), non-alcoholic steatohepatitis (n=17), primary sclerosing cholangitis (n=4).
If you wish to use the HepaFat-Scan service or would like to know more about HepaFat-Scan, please send an email to [email protected] for further information.
HepaFat-Scan Resources
Click the link(s) below to access our resources. Need something else? Please contact our team at [email protected] for further information.